Best Practices for Engineering ML Pipelines - Part 2
Posted on Mon 07 November 2022 in machine-learning-engineering • Tagged with python, machine-learning, mlops, kubernetes, bodywork

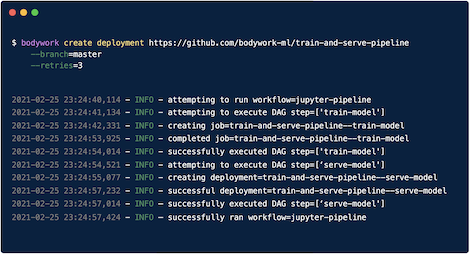
This is the second part in a series of articles demonstrating best practices for engineering ML pipelines and deploying them to production. In the first part we focused on project setup - everything from codebase structure to configuring a CI/CD pipeline and making an initial deployment of a skeleton pipeline …
Continue reading